I am a structural biologist interested in understanding the molecular principles of protein folding and misfolding. How cells manage partially folded and misfolded proteins and the effect of protein misfolding on protein quality control machineries of the cell are some of the questions that I am excited to investigate.
Below are some of my current and past projects:
Molecular mechanism of co-translational mRNA surveillance by the ER stress sensor, IRE1
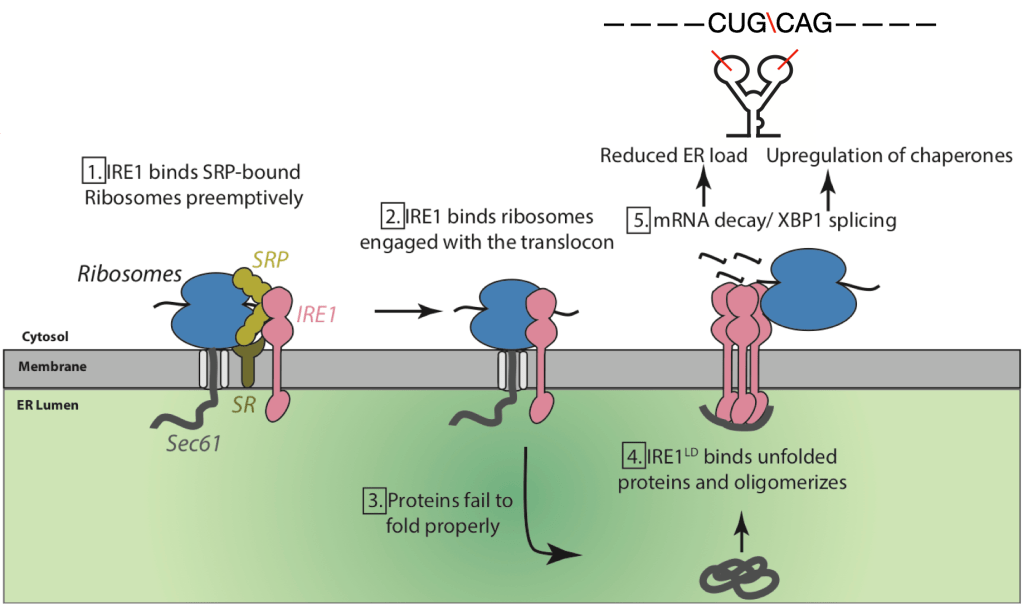
My current project focuses on the molecular mechanisms of ER quality control and its maladaptation under stress. Currently, I am working on IRE1 (Inositol Requiring Enzyme1) which acts as a gatekeeper of the ER. It is a kinase/RNase which gets activated upon accumulation of unfolded proteins in the ER and activates a signaling network by cleaving ER-targeted mRNAs to correct the defect. How IRE1 recognizes its mRNA substrates has been unknown. Recently, the Walter lab discovered that IRE1 binds the co-translational targeting machinery composed of ribosomes, Sec61 and SRP (Acosta-Alvear et al., 2018). This previously unseen interaction suggests that IRE1 monitors and exerts control on the translational status of ER-targeted proteins co-translationally. I have focused on gaining a molecular and functional understanding of this complex. I isolated IRE1-ribosome complexes and determined their architecture using single-particle cryoEM. To trap these transient complexes, I utilized specialized microscopy grids developed in the lab of my co- mentor, Prof. David Agard (Palovcak et al., 2018; Wang et al., 2020). The structures revealed that IRE1 associates with a subset of ER-targeted ribosomes in a supramolecular complex that also includes the ER membrane channel, Sec61 and surveys the mRNA co-translationally. In complementary efforts, I used deep sequencing to identify the mRNAs associated by IRE1 and discovered that IRE1-bound ribosomes are enriched in mRNAs with putative cleavage sites. Together my structural and functional studies suggest that monomeric IRE1 serves as a sensor on the mRNA entry site of ribosomes primed to cleave mRNAs upon activation (Sangwan et al. in preparation).
Translational regulation under stress

As cells experience stressors such as amino acid deprivation, viral infections and ER stress, they shut down global translation while maintaining the pool of essential proteins such as transcription factors, and chaperones. To continue the translation of these select transcripts, they utilize novel proteins and features in the mRNA sequence itself. One such protein is eukaryotic initiation factor 2A (eIF2A). eIF2A is a putative alternate translation initiation factor involved in translation of specific mRNAs possibly utilizing alternate start codons under stress. Its specific role and mechanism of action has been completely unknown. I am investigating the role of eIF2A in translation using single particle cryoEM and functional assays. My work suggests that eIF2A binds at a discrete site on the 40S ribosomal unit utilizing its disordered regions to connect with the 40S (Sangwan et al. in preparation).
Design of aerosolized spray against SARS-CoV2 virus

Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) virus enters host cells via an interaction between its Spike protein and the host cell receptor angiotensin-converting enzyme 2 (ACE2). I developed nanobodies that disrupt the interaction between Spike and ACE2 by screening a yeast surface-displayed library of synthetic nanobody sequences. I then used cryo-EM to obtain the atomic level details of the nanobody. One nanobody, Nb6, binds Spike in a fully inactive conformation with its receptor binding domains locked into their inaccessible down state, incapable of binding ACE2. I utilizied biochemical assays such as affinity maturation and structure-guided design of multivalency to yield a trivalent nanobody, mNb6-tri, with femtomolar affinity for Spike and picomolar neutralization of SARS-CoV-2 infection. mNb6-tri retains function after aerosolization, lyophilization, and heat treatment, which enables aerosol-mediated delivery of this potent neutralizer directly to the airway epithelia.
Structure-guided inhibitors of alpha-synuclein aggregation in Synucleinopathies

α-Syn is a protein that forms amyloid fibrils in Parkinson’s disease and several related diseases together known as Synucleinopathies. Intriguingly, it is known that pathology progresses through interconnected brain regions a phenomenon known as seeding where preformed small fibrils can template the formation of fibrils. A short segment (residues 68-78) is known to be necessary for full-length α-syn aggregation both in vitro and in vivo. As a collaborative effort, I determined the atomic structure of a short segment known to be necessary for full-length α-syn aggregation both in vitro and in vivo (Rodriguez et al. Nature 2015). I then utilized the structure to design a line of peptide inhibitors that prevent α-syn aggregation and seeding (patent filed 2015, Sangwan et al. eLife 2020). The inhibitors bind with high affinity (Kd ~ 0.5 μM) to α-syn fibrils and prevent seeded propagation of α-syn aggregates. My approach displayed the potential of rational structure-based drug design to halt protein aggregation.
Molecular mechanism of SOD1-linked cytotoxicity in ALS

Fibrils and oligomers are the aggregated protein agents of neuronal dysfunction in ALS diseases. Whereas we now know much about fibril architecture, atomic structures of disease-related oligomers have eluded determination. I determined the structure of a short segment of SOD1 by X-ray crystallography. The segment adopts a corkscrew-like fold forming oligomers. I then made mutations in both the segment and in full-length protein that disrupt corkscrew formation. I found that mutations that prevent formation of this structure eliminate cytotoxicity of the segment in isolation as well as cytotoxicity of the ALS-linked mutants of SOD1 in primary motor neurons and in a zebrafish model of ALS. I also determined 3 crystal structures of overlapping segments from SOD1. The crystal structures reveal three different architectures: corkscrew oligomeric structure, non-twisting curved sheet structure and a steric zipper proto‐filament structure. My work highlights the polymorphism of the segment 28–38 of SOD1 and identifies the molecular features of amyloidogenic entities.